Colon adenocarcinoma histological grading based on multivariable analysis of the status of tumor cell nuclear apparatus
Grabovoy A.N.1, Antoniak S.N.2, Savchyn T.M.
- 1National Cancer Institute, Kyiv
- 2Institute of Epidemiology and Infectious Disease named after L.V. Gromashevskogo
Summary. The aim of the work: development of a model of histological grading of colon adenocarcinoma (AC) based on multivariable logistic regression analysis of the status of tumor cell nuclear apparatus. In the current study, biopsy material and material obtained during surgical treatment of patients with colon AC or benign colon neoplasms were analyzed using histological, morphometric, densitometric, immunohistochemical methods and mathematical modelling. To improve separation of tumors of adjacent grades different models with different coefficients and different set of parameters are required. Thus, the key parameters for classifying benign neoplasms from AC are: average DNA content, proportion of viable cancer cells, number of nucleolar organizers, proliferation index, and expression level of p53. The Model for separating AC G1 from G2 is characterized by a similar set of parameters and contains the average size of a nucleolar organizer regions as an additional parameter. The difference between AC G2 and G3 is more vague, and these histological types are distinguished by only two features, by Bcl-2 and p53 tumor expression level. The proposed model for grade estimating for patients with colon AC demonstrated good quality.
Submitted 25.01.2018
Received 28.02.2018
Introduction
The differentiation tumor grade (G) nowadays is the main histological prognostic criterion for colorectal cancer prognosis [6, 7] and is an essential component of the pathogistological conclusion in routine clinical practice. But accepted methods of tumor differentiation are often biased, since it is difficult to avoid human factor, assess the tissue sample rigorously and, therefore, make a correct treatment choice. Recently obtained data has shows that the predictive significance of the tumor grade strongly depends on the microsatellite instability (MSI) status. It has been shown that cases with high-grade colorectal tumors have a poor prognosis in the case of microsatellite stability (MMS). And in the case of high MSI (MSI-H), they behaves as highly differentiated tumors [13]. There is also an evidence that the identifying of low-differentiated colon adenocarcinoma (AC) clusters is more reproducible and has a higher predictive value in comparison to the grade system [2].
In addition, a large number of different tumor features were presented as independent prognostic criteria of AC [3, 10]. They are: amount of DNA, mitotic activity, status of adhesive complexes and receptors, etc. However, none of them has been accepted as a significant criterion. This is the result of the heterogeneity of colorectal AC [12] in consequence of various pathogenetic mechanisms of tumor development, the diversity of genetic damage and involvement of several epigenetic mechanisms [1, 4, 8, 15].
The aim of the work: development of the test (system) to improve the tumor differentiation based on a multivariable analysis of the status of tumor cell nuclear apparatus in a patients with colorectal AC.
Material and methods
The study was performed on 127 samples of biopsy material or material obtained during surgical treatment of patients with colon AC or benign colon neoplasms: conventionally normal colon tissue (C; n=10); polyps and adenomas (В; n=15); AC of different differentiation grades: G1 (n=27); G2 (n= 62); G3 (n=13).
The material was fixed in a buffered 10% formalin (pH 7.4, 4 °С, 24 h) and embedded in paraplast with the use of Histos-5 histoprocessor (Milestone, Italy). From paraffin blocks, 5 µm sections were cut using Microm НМ325 (Thermo Scientific, Germany). The sections were stained with hematoxylin and eosin for common tumor assessment, or with Einarson’s gallocyanin chrome alum stain (pH 1.62, 37 °С, 24 h) for determination of nucleic acid content [11, 14]. In each case, a part of sections was treated with RNAase (MACHEREY-NAGEL GmbH & Co. KG, Germany) for RNA destruction [11]. Nucleolar organizers (NOR) were detected using silver nitrate impregnation. Immunohistochemical reactions were performed with the use of mouse monoclonal antibodies against human Ki-67 (MIB-1, Dako, Denmark), human Bcl-2 oncoprotein (Dako, Denmark), and human р53 protein (Dako, Denmark) according to the instructions of the manufacturer and visualized using detection system EnVisionTM FLEX (Dako, Denmark). The sections were additionally stained with Gill hematoxylin.
The preparations were examined and photographed using microscope Nikon Eclipse 80i supplied with camera DS-5SMc/L2 at standardized conditions, images (×400, 1280×960 pixels RGB) were processed using analysis system ImageJ 1,46. In images of the preparations stained with gallocyanin chrome alums, there have been determined cross-sectional area of cell nucleus, medium (DM) and integral optical density of nucleus, and the quantity of nuclear DNA (NDNA) in each 30 tumor cells. For calculation of the latter index, DNA content in lymphocyte nuclei was accepted as a unit. This population of cells was divided into three morphofunctional types of cells using the original method based on relative DNA content and area of nucleus. These types were as follows: conventionally viable cells with “normal” nucleus, cells with pyknotic nucleus and cells with lytic nucleus. For each case, relative numbers of these three types of cells were evaluated: Nn, Np, Nl, respectively [5].
The tumors were distributed into two subgroups by average DNA content: D — average DNA content up to 1.2, P — average DNA content higher than 1.2.
In silver-impregnated sections, in each of 50 tumor cells there was determined cross-sectional area of nucleus, number of NOR (nNOR) in nucleus, and diameter of each NOR with following calculation of their total volume (vNOR). In each tumor, the percentage of cells expressing Ki-67, Bcl-2, and р53 has been determined. The percentage of reaction-positive cells was calculated in 5 different vision fields at 400× magnification.
Statistical analysis was performed using IBM SPSS Statistics 15. One sample Kolmogorov — Smirnov test was applied for assessment of normality. Correlation analysis was conducted by Spearman rank correlation. Assessments of differences between groups were carried out using Mann — Whitney U test.
Logistic regression was applied as a classification tool. The generalized equation of regression was hypothesized as follows:
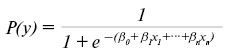 ,
,
where: P(y) — the probability of an event y;
β0 — a constant;
βn — regression parameters;
xn — features of nuclear apparatus of tumor cell.
As a basis, the hierarchical classification method was used. It involves sequential separation of a set of objects into a subset and, reminds a tree structure. The main criterion for the separation is a certain feature that describes the difference of a data subset the most. But in our case, the main criterion is not a discrete feature, but a set of different features transformed by logistic form into binary form [9]. On the first step, the tumor set are divided into benign and malignant neoplasms (Model I). On the next step, the malignant tumors is divided into well-differentiated AC (G1) and AC moderately (G2) and low-differentiated (G3) (Model II). On the last stage, AC G2 and AC G3 are finally divided (Model III).
For assessing the quality of obtained models, they were compared with the null model using likelihood ratio test (LR). Moreover, the quality of obtained models was assessed by correct classification rate and ROC-analysis. The area under curve (AUC) points on the quality of model: 0.5–0.6 — fail, 0.6–0.7 — poor, 0.7–0.8 — fair, 0.8–0.9 — good, 0.9–1 — excellent. p-values lower or equal to 0.05 were considered as statistically significant.
Results and discussion
On the first step of the study quantitative values of the features of the cell nuclei were established for AC of different grades, adenomas and for epithelium of a normal colon (Table 1). For each feature pairwise comparison was applied for study groups (Table 2).
| Feature | Conventionally normal colon tissue | Benign colon neoplasms | Colon AC | ||
| G1 | G2 | G3 | |||
| NDNA | 1,30±0,07 | 2,31±0,31 | 2,39±0,21 | 1,98±0,08 | 2,06±0,16 |
| nNOR | 1,70±0,08 | 3,04±0,14 | 2,87±0,17 | 2,37±0,10 | 2,25±0,19 |
| vNOR | 1,96±0,09 | 2,80±0,19 | 2,62±0,19 | 2,86±0,16 | 3,25±0,43 |
| v/NOR | 1,15±0,04 | 0,92±0,04 | 0,92±0,06 | 1,30±0,09 | 1,56±0,28 |
| Ki-67, % | 10,22±3,49 | 34,56±6,37 | 37,19±5,15 | 45,93±3,39 | 49,09±8,50 |
| Bcl-2, % |
85,52±4,02 | 36,13±10,70 | 42,76±8,43 | 41,21±5,57 | 17,60±7,48 |
| p53, % | 32,21±3,82 | 15,95±8,05 | 52,21±7,60 | 69,81±4,39 | 37,43±10,21 |
| Nn, % | 58,68±6,56 | 69,16±7,26 | 81,12±3,27 | 84,70±1,31 | 86,67±4,03 |
| Nl, % | 0,00 | 8,44±5,34 | 7,59±2,02 | 7,64±0,93 | 5,38±1,28 |
| Np, % | 41,32±6,56 | 22,40±6,62 | 11,29±3,42 | 7,66±1,34 | 7,95±3,90 |
NDNA — content of DNA in nucleus; nNОR — number of nucleolar organizers; vNОR — total volume of nucleolar organizers; v/NOR — average volume of nucleolar organizers; Ki-67, Bcl-2, p53 — expression level of corresponding marker in tumor (%); Nn, Nl, Np — proportion of conventionally viable cells with «normal» nucleus, cells with lytic nucleus and cells with pyknotic nucleus, respectively (%).
| Feature | Comparing groups | ||
| K-B/G1-2-3 | G1/G2-3 | G2/G3 | |
| NDNA | <0,01 | =0,171 | =0,828 |
| nNOR | =0,946 | =0,004 | =0,700 |
| vNOR | =0,073 | =0,416 | =0,308 |
| v/NOR | =0,379 | =0,001 | =0,338 |
| Ki-67, % | <0,01 | =0,075 | =0,651 |
| Bcl-2, % | =0,123 | =0,738 | =0,304 |
| p53, % | <0,01 | =0,131 | =0,017 |
| Nn | <0,01 | =0,412 | =0,133 |
| Nl | <0,01 | =0,448 | =0,527 |
| Np | <0,01 | =0,326 | =0,431 |
NDNA — content of DNA in nucleus; nNОR — number of nucleolar organizers; vNОR — total volume of nucleolar organizers; v/NOR — average volume of nucleolar organizers; Ki-67, Bcl-2, p53 — expression level of corresponding marker in tumor (%); Nn, Nl, Np — proportion of conventionally viable cells with «normal» nucleus, cells with lytic nucleus and cells with pyknotic nucleus respectively (%); bold values indicate significant difference (p<0.05).
Statistically significant results were obtained for Model I (identifying K+B/G1+G2+G3): NDNA, Ki-67 and p53 expression, Nn, Nl and Np (see Table 2). However, the possible effect of indicators interaction was not rejected. One of the tested models took into account insignificant features (nNOR), and their moderation effect with others (NDNA and nNOR, for example). As a result, a statistically significant model was obtained (Table 3).
| Feature | β | SE | p |
| Model I (benign and malignant) | |||
| NDNA | 5,406 | 1,967 | <0,01 |
| nNOR | 3,195 | 1,329 | =0,016 |
| NDNA*nNOR | −1,864 | 0,659 | <0,01 |
| Ki-67, % | 0,030 | 0,014 | =0,028 |
| p53, % | 0,079 | 0,021 | <0,01 |
| p53(+/−) | 3,478 | 1,234 | <0,01 |
| Nn | 0,058 | 0,018 | <0,01 |
| Constant | −17,291 | 4,572 | <0,01 |
| Model II (G1/G2–3) | |||
| NDNA | −1,968 | 0,587 | =0,001 |
| nNOR/Ki-67 | −0,090 | 0,038 | =0,018 |
| NDNA*nNOR/Ki-67 | 0,034 | 0,016 | =0,034 |
| v/NOR | 2,199 | 0,888 | =0,013 |
| p53, % | 0,011 | 0,007 | =0,094 |
| D/P | −2,503 | 0,951 | =0,008 |
| Constant | 2,901 | 1,481 | =0,050 |
| Model ІІІ (G2/G3) | |||
| p53, % | −0,040 | 0,018 | =0,026 |
| Bcl-2, % | −0,031 | 0,014 | =0,030 |
| Bcl-2(+/–) | −1,674 | 0,857 | =0,051 |
| p53(+/–) | −2,275 | 1,408 | =0,106 |
| Constant | 2,684 | 1,456 | =0,065 |
NDNA — content of DNA in nucleus; nNОR — number of nucleolar organizers; v/NOR — average volume of nucleolar organizers; Ki-67, Bcl-2, p53 — expression level of corresponding marker in tumor (%); p53(+/−), Bcl-2(+/−) — positive or negative expression of corresponding marker; Nn — proportion of conventionally viable cells with «normal» nucleus (%); β — coefficient; SE — standard error of coefficient; p — significance.
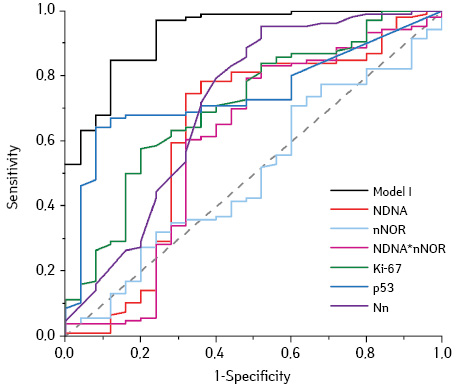
And this model in addition to common independent parameters (Ki-67 (%), p53 (%), Nn) contains the interaction effect of NDNA and nNOR (Table 3). The appearance of nNOR in the equation is related with the combined effect of this parameter and NDNA, since polyploidy shows an increase and the nNOR. Positive values of the coefficients (β) indicate that effects of features are related to the probability of classify the carcinoma. In other words increasing of NDNA, nNOR, Ki-67, Nn, p53 occurs during malignant transformation. The correct classification rate for Model I was 91.6%. The obtained value of AUC for Model I was 0.930 (Fig. 1), indicating that model quality as for diagnostic test is excellent. Note that the value of the AUC for the whole model was greater than the AUC for individual parameters. It displays the combined effect of the features, and indicates stand alone using is unsuitable.
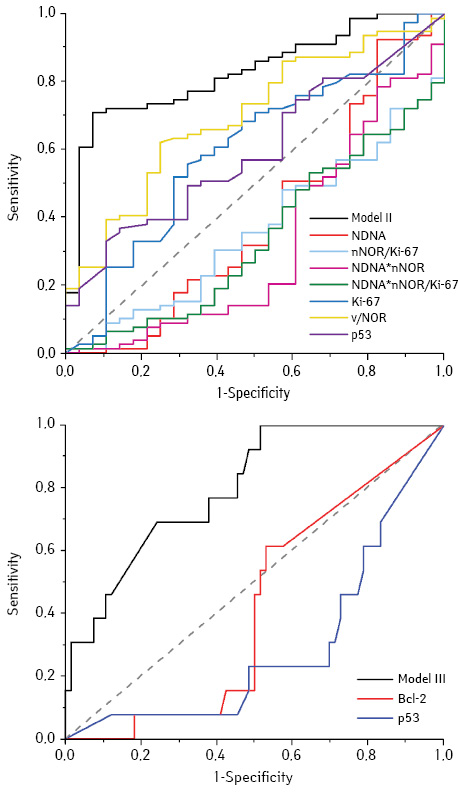
Models II and III were built to separate degree of differentiation (G) of AC (Table 3). Model II segregates G1 from G2 and G3. After that Model III separetes G3 from G2. Each of these models, consists of diverse set of quantitative and qualitative features. Accordingly, DNA, nNOR, vNOR, Ki-67 (%), p53 (%), and tumor polyploidy (D, P) comprises into Model II. It can believed that these features could be indirectly related to the different mechanisms of development, for example, aneu-/polyploidy is usually a tag of chromosomal instability [4, 8]. The negative value of β for polyploidy indicates that increasing of this feature is significant for G2 and G3. The correct classification rate for Model II was 75.7%.
For Model III, the set of significant parameters was the poorest, indicating a very large similarity between G2 and G3 groups. The correct classification rate for Model III was 87.3%. Tumor expression rate (%) of Bcl-2 and p53 plays a key role in the separation of these grades of AC. Besides, an additional parameters — the tumor phenotypes based on expression of p53 and Bcl-2 (p53(+/–), Bcl-2(+/–)) were included to improve significance of p53 (%) and Bcl-2 (%).
The quality assessment of Models II and III using ROC analysis (Fig. 2, Table 4) showed that the AUC values for both models are higher than for individual parameters. It is 0.816 and 0.801, respectively, which indicates a very good quality of the models.
| Feature | AUC | SE | p | CI (95%) | |
| Model I (benign and malignant) | |||||
| Model І | 0,930 | 0,028 | <0,01 | 0,876 | 0,985 |
| NDNA | 0,634 | 0,074 | =0,038 | 0,489 | 0,778 |
| nNOR | 0,499 | 0,064 | =0,993 | 0,374 | 0,625 |
| NDNA*nNOR | 0,598 | 0,074 | =0,126 | 0,453 | 0,744 |
| Ki-67, % | 0,711 | 0,058 | =0,001 | 0,598 | 0,824 |
| p53, % | 0,744 | 0,048 | <0,01 | 0,650 | 0,837 |
| Nn | 0,715 | 0,067 | =0,001 | 0,584 | 0,846 |
| Model ІІ (G1/G2–3) | |||||
| Model ІІ | 0,827 | 0,042 | =0,000 | 0,744 | 0,910 |
| NDNA | 0,396 | 0,068 | =0,105 | 0,263 | 0,530 |
| nNOR/Ki-67 | 0,362 | 0,058 | =0,031 | 0,249 | 0,475 |
| NDNA*nNOR | 0,323 | 0,064 | =0,005 | 0,198 | 0,448 |
| NDNA*nNOR/Ki-67 | 0,339 | 0,058 | =0,012 | 0,225 | 0,453 |
| Ki-67 | 0,598 | 0,063 | =0,123 | 0,475 | 0,722 |
| v/NOR | 0,698 | 0,055 | =0,002 | 0,590 | 0,806 |
| p53, % | 0,596 | 0,058 | =0,133 | 0,481 | 0,710 |
| Model ІІІ (G2/G3) | |||||
| Model ІІІ | 0,801 | 0,060 | =0,001 | 0,684 | 0,919 |
| Bcl-2, % | 0,417 | 0,072 | =0,344 | 0,443 | 0,724 |
| p53, % | 0,290 | 0,076 | =0,017 | 0,562 | 0,858 |
NDNA — content of DNA in nucleus; nNОR — number of nucleolar organizers; v/NOR — average volume of nucleolar organizers; Ki-67, Bcl-2, p53 — expression level of corresponding marker in tumor (%); Nn — proportion of conventionally viable cells with «normal» nucleus (%); SE — standard error of coefficient; p — significance; CI — confidence interval.
Since there is no general-purpose approach for specificity and sensitivity optimization, researcher tunes the model according to a particular case. For example, model classifies G3 with high sensitivity, but in contrast separates G1 from G2 and G3 with a smaller sensitivity. This, in turn causes certain corrective effect on the quality of the models.
In this study model optimization was based on the statement that the idealized model demonstrates 100% sensitivity and specificity. Thus, an optimal value of sensitivity and specificity determination was conducted with varying the value of the cutoff point of quantifier (from 0 to 1). For this purpose, the sensitivity curves and specificity curves for each model were plotted as a function of the cut-off values of quantifier (Fig. 3). At the intersections of the sensitivity and specificity curves (points A, B, and C), values are equal (Se = Sp). One can see from the plot that points for Models I and Model II lie closer to each other (points A and B, respectively) than for Model III (point B). Points B and C are approximately at the same level of sensitivity and specificity — 70%. Point A lies a bit higher — 85%.
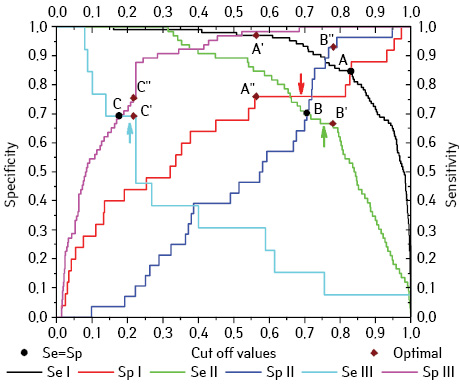
Next to point C (intersection of the curves of Se III and Sp III) (see Fig. 3) there is a “plateau” of sensitivity (Fig. 3, turquoise arrow). For this line segment, the sensitivity remains at the same level, but the specificity is going to increase during the shifting the cut-off threshold to the right up to 0.216 (points C’ and C”). This effect allows us to increase the specificity from 69.7% to 75.8% without loss of sensitivity. For point B, situation is very similar (Fig. 3, green arrow), therefore, the cut-off threshold shifting causes minimal sensitivity loss with significant incrementation of specificity of Model II.
For Model I, there is also a large enough plateau (Fig. 3, red arrow). Therefore, minimal reduction of specificity from 85 to 76%, increases the sensitivity of the Model I from 85 to 97.2% (points A’ and A”) with the cut-off of 0.56. Note that this cut-off value satisfies the Se + Sp = max criterion. As a result, approximately the same values of sensitivity and specificity were obtained for three models. The sensitivity and specificity values lie in the range of 70–75%.
Correct classification rate for optimized Model I was 93.1%. Three from 106 cases with AC were assigned to the benign neoplasms improperly (Fig. 4). All three samples represent the absence of expression of p53, low values of the Ki-67 index (20; 8, and 43%). For DNA content and the number of nucleoli, their values did not act as a significant criterion for good or malignant neoplasms differentiation. Six cases from 25 benign neoplasms were attributed to AC. High values of Nn (%) (from 77 to 97%) is a reason of that effect. One of the cases, represents high expression of p53 (93%), high DNA content (NDNA=3) and a large number of nucleoli (nNOR=4).
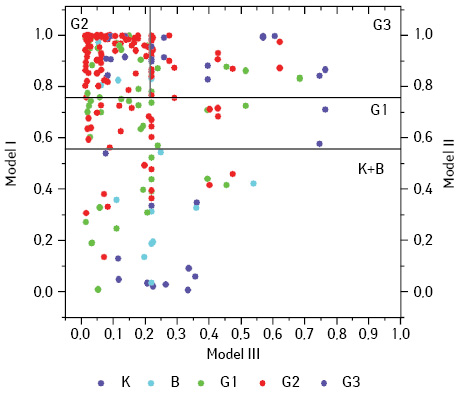
For optimized Model II with 0.76 cutoff point, correct classification rate was 76.6%. Three cases from 28 with G1 were assigned to higher AC grades (see Fig. 4). Two of them were characterized by increased vNOR. The third case had a relatively normal value of vNOR, but a very high level of expression of p53 (90%). 22 cases from 79 that have been identified as G2–3 have now had been classified G1. These ACs represents large variability of the characteristics, it makes impossible to classify them correctly.
Model III now shows correct classification rate in 74.7%. Four from 13 cases of AC G3 have been classified as G2 by optimized model (see Fig. 4). These cases represented high levels of p53 expression and Bcl-2. However, 16 from 66 cases with G2 were classified as G3, all the cases represents low values of the above parameters.
Thus, the conducted study has shown that ACs are characterized by a large variety of status of nucleus, proliferative activity and expression of factors regulating apoptosis. In our opinion, it is associated with different pathogenetic mechanisms involved in colorectal cancer development [1, 3, 4, 15]. Based on our previous research [6], and [3, 10, 12], it can be assumed that separate features have a weak, if any, link with the colorectal cancer prognosis. It negates their use as prognostic criteria. However, using some of the key features of status of the nuclei of epithelial tumors of the colon and/or their derivatives in a multivariate approach allows us to increase the reliability of the determination of the AC grade due to the objectification of this procedure.
The great variability of the nuclei of AС G2 cells and significant dispersion in the prognosis are serious arguments to change a traditional three-level histological grading to two-level grading for this type of cancer [2]. Moreover a very tiny set of significant features can be used to distinguish AC G2 from AC G3.
It is worthwhile to say that the presence of different various mechanisms of AC development has a significant effect on the values of these features in the personalized assessment of AC. Therefore, the problem of an objective assessment of histological grade at this stage remains unresolved. In prospect to find significant parameters that will allow separation with a clinically acceptable level of accuracy, taking into account all the difficulties of the colorectal cancer development.
Conclusions
AC of the colon of different degrees of differentiation are quite stand in marked contrast from each other by a set of features of the status of the tumor cell nucleus. Objective determination of AC grade using such features as the amount of DNA, the state of NOR, the tumor expression level of Ki-67, p53 and Bcl-2 as independent criteria is unsuitable due to high variability of these features. On the contrary using these features and their derivatives in multivariate approach based on logistic classifier dramatically increases the quality of determining the grade of AC of the colon. To improve separation of tumors of adjacent grades different models with different coefficients and different sets of parameters are required. Thus, the key parameters for classifying benign neoplasms from AC are: average DNA content, proportion of viable cancer cells, nNOR, proliferation index, and expression level of p53. Model for separating AC G1 from G2 A is characterized by a similar set of parameters and contains the average size of a NOR regions as an additional parameter. The difference between AC G2 and G3 is more vague, and these histological types are distinguished by only two features, by Bcl-2 and p53 tumor expression level.
References
1. Bardhan K., Liu K. (2013) Epigenetics and Colorectal Cancer Pathogenesis. Cancers, 5(2): 676–713.
2. Barresi V., Reggiani L.B., Ieni A. et al. (2015) Histological grading in colorectal cancer: new insights and perspectives. Histol. Histopathol., 30(9): 1059–1067.
3. Coppedè F., Lopomo A., Spisni R., Migliore L. (2014) Genetic and epigenetic biomarkers for diagnosis, prognosis and treatment of colorectal cancer. World J. Gastroenterol., 20: 943–956.
4. Davoli T., Lang T. (2011) The Causes and Consequences of Polyploidy in Normal Development and Cancer. Ann. Rev. Cell Dev. Biol., 27: 585–610.
5. Grabovoy A.N., Antoniuk S.A., Silchenko V.P. et al. (2015) Pytomyy vmist DNK u yadrakh klityn i klitynna heterohennistʹ adenokartsynom tovstoyi kyshky [Specific cell’s nuclei DNA content and cellular heterogeneity of colon adenocarcinoma]. Pathologia, 1(33): 44–47.
6. Grabovoy A.N., Kolesnik O.O., Savchyn T.M., Antoniuk S.A. (2016) Modelling of survival of patients with colon adenocarcinoma based on multivariable analysis of the state of cancer cell nuclear apparatus. Exp. Oncol., 38 (1): 45–48.
7. Hamilton S.R., Bosman F.T., Boffetta P. et al. (2010) Carcinoma of the colon and rectum. In: WHO classification of tumors of the digestive system. Bosman T., Carneiro F., Hruban R.H. and Theise N.D. (eds). IARC Press, Lyon: 134–146.
8. Holland A., Cleveland D. (2012) Losing balance: the origin and impact of aneuploidy in cancer. EMBO, 13: 501–514.
9. Irwin R.J., Irwin T.C. (2011) A principled approach to setting optimal diagnostic thresholds: where ROC and indifference curves meet. Eur. J. Intern. Med., 22(3): 230–234.
10. Kulendran M., Stebbing J.F., Marks C.G., Rockall T.A. (2011) Predictive and prognostic factors in colorectal cancer: a personalized approach. Cancers (Basel), 29: 1622–1638.
11. Luppa Kh. (1980) Osnovy hystokhymyy (Per. s nemets.) [Basic histochemistry (Tran. from Germ.)]. Myr, Moscow, 344 p.
12. Marusyk A., Almendro V., Polyak K. (2012) Intra-tumour heterogeneity: a looking glass for cancer? Nat. Rev. Cancer, 12(5): 323–334.
13. Rosty C., Williamson E.J., Clendenning M. et al. (2014) Should the grading of colorectal adenocarcinoma include microsatellite instability status? Hum. Pathol., 45(10): 2077–2084.
14. Tashke K. (1980) Vvedenye v kolychestvennuiu tsyto-hystolohycheskuiu morfolohyiu. (Per. s rum.) [Introduction to quantitative cyto-histological morphology. (Trans. from Roman.)]. Yz. Akad. Sots. Resp. Rumynyi., Moscow, 192 p.
15. Zoratto F., Rossi L., Verrico M. et al. (2014) Focus on genetic and epigenetic events of colorectal cancer pathogenesis: implications for molecular diagnosis. Tumour Biol., 35 (7): 6195–6206.
Grabovoy Alexandr
33/43 Lomonosova str., Kyiv 03022
National Cancer Institute
E-mail: angrabovoy@gmail.com
Адреса:
Грабовий Олександр Миколайович
03022, Київ, вул. Ломоносова, 33/43
Національний інституту раку
E-mail: angrabovoy@gmail.com














Leave a comment